
Vegetative compatibility and parasexual segregation in Colletotrichum lindemuthianum, a fungal pathogen of the common bean M.A.A. Castro-Prado, C.B. Querol, J.R. Sant’Anna, C.T. Miyamoto,C.C.S. Franco, C.A. Mangolin and M.F.P.S. Machado Genet. Mol. Res. 6 (3): 634-642 (2007) ABSTRACT. The heterokaryotic and vegetative diploid phases of Colletotrichum lindemuthianum are described using nutritional and biochemical markers. Nitrate non-utilizing mutants (nit), derived from R2047, R89, R73, R65, and R23 isolates, were paired in all possible combinations to obtain heterokaryons. Although pairings R2047/R89, R2047/R73, R65/R73, and R73/R23 showed complete vegetative incompatibility, prototrophic heterokaryons were obtained from pairings R2047/R65, R2047/R23, R65/R89, R65/R23, R73/R89, R89/R23, R2047/R2047, R65/R65, R89/R89, R73/R73, and R23/R23. Heterokaryons gave rise to spontaneous mitotic segregants which carried markers corresponding to one or the other of the parental strains. Heterokaryons spontaneously produced prototrophic fast-growing sectors too, characterized as diploid segregants. Diploids would be expected to yield auxotrophic segregants following haploidization in basal medium or in the presence of benomyl. Parental haploid segregants were in fact recovered from diploid colonies growing in basal medium and basal medium containing the haploidizing agent. Although barriers to the formation of heterokaryons in some crosses were detected, the results demonstrate the occurrence of parasexuality among vegetative compatible mutants of C. lindemuthianum. Key words: Colletotrichum lindemuthianum, Heterokaryosis, Esterase isoenzymes, Nitrate non-utilizing mutants, Vegetative compatibility groups INTRODUCTION Colletotrichum lindemuthianum (teleomorph Glomerella lindemuthiana) is the causal agent of anthracnose, one of the main diseases of the common bean (Phaseolus vulgaris). Anthracnose occurs principally in leaf veins and pods, with almost total production loss (Jerba et al., 2005). Although the sexual phase of C. lindemuthianum has already been reported in the laboratory, sexual reproduction in field conditions is not common. However, the pathogen shows high genetic variability which interferes with the cultivars’ resistance durability, with subsequent difficulty in the effective control of anthracnose (Kimati and Galli, 1970; Balardin et al., 1997; Mahuku and Riascos, 2004). The formation of conidial anastomosis tubes between conidia within acervuli has been observed during C. lindemuthianum conidiogenesis and described as a mechanism that favors exchange of nuclear material and organelles between incompatible strains. Conidial anastomosis tubes were detected in pairs or in multiple associations of conidia (Roca et al., 2003). Hyphal anastomoses among vegetative compatible isolates also favor exchange of genetic material in filamentous fungi and lead to the formation of vegetative heterokaryons (i.e., cells containing two genetically different nuclei), which have been characterized as the first stage of the parasexual cycle (Pontecorvo et al., 1953; Saupe, 2000). Haploid nuclei in the heterokaryotic mycelium fuse together and form diploid nuclei. Through successive processes of chromosome non-disjunction, the latter give rise to paternal haploids or recombinants (Pontecorvo et al., 1953; Miyamoto et al., 2007). The viability of heterokaryons is regulated by multiple vegetative incompatibility loci, named vic (for vegetative incompatibility) or het (for heterokaryon incompatibility), so that only compatible strains, believed to be clonally related, may produce stable heterokaryons among themselves (Saupe, 2000). Actually, nitrate non-utilizing mutants (nit) of Magnaphorte grisea and Fusarium species have been used to force heterokaryons between nearly isogenic strains, and to test isolates for vegetative compatibility (Klittich and Leslie, 1988; Baayen et al., 1998; Correl et al., 2000). The aim of the present study was to obtain nit mutants from C. lindemuthianum isolates and to use these auxotrophic mutants to obtain heterokaryons and parasexual segregants. Whereas nitrate non-utilizing mutants were used to form heterokaryons and diploids, native polyacrylamide gel electrophoresis was used to show the differential expression of α- and β-esterase in haploid and diploid strains of C. lindemuthianum isolates, as an additional marker. MATERIAL AND METHODS StrainsRaces R2047 and R23 of C. lindemuthianum were kindly donated by Dr. Maria Celeste Vidigal (Departamento de Agronomia, Universidade Estadual de Maringá, Paraná, Brazil), while races R89, R73 and R65 were obtained from the Embrapa Arroz e Feijão, Goiás, Brazil. Culture stocks were maintained at 5ºC in amber flasks with basal medium (BM). MediaBM was prepared with 1000 mL distilled H2O, 30 g saccharose, 1 g KH2PO4, 0.5 g MgSO2. 7 H2O, 0.5 g KCl, 10 mg FeSO4 . 7 H2O, 15 g agar, and 0.2 mL trace element solution (95 mL distilled H2O, 5 g citric acid, 1 g Fe(NH4)2(SO4)2 . 6 H2O, 0.25 g CuSO4 . 5 H2O, 50 mg MnSO4 . 5 H2O, 50 mg H3BO3, and 50 mg Na2MoO2 . H2O). Minimal medium (MM) consisted of Czapek-Dox with 1% (w/v) glucose and 15 g agar. Complete medium (CM) consisted of 1000 mL distilled H2O, 10 g glucose, 2 g peptone, 0.5 g yeast extract, 1 g hydrolyzed casein, 4 mg inositol, 2 mg choline chloride, 2 mg pantothenic acid, 1 mg nicotinic acid, 1 mg riboflavin, 0.1 mg 4-aminobenzoic acid, 0.5 mg folic acid, 0.5 mg pyridoxine, 0.2 mg thiamine, and 2 µg biotin added to MM. Isolation of mutantsMutants that were unable to use nitrate as a nitrogen source (nit) were obtained in BM + NaNO3 (0.2%) + KClO3 (3.0%). Mycelium plugs 5 mm in diameter were removed from the periphery of C. lindemuthianum colonies cultivated in CM and transferred to the center of petri dishes with BM + NaNO3 + KClO3. Dishes were incubated at 22ºC for 25 days. Chlorate-resistant mutants were then identified, isolated and transferred to petri dishes with BM, supplemented with different nitrogen sources: BM + sodium nitrate (0.2%), BM + potassium nitrite (0.085%), BM + hypoxanthine (0.01%), and BM + ammonium tartrate (0.092%). Plates were incubated at 22ºC for 12 days, and mutants were phenotypically characterized according to Klittich and Leslie (1988). Hyphal anastomosesComplementary nit mutants from the isolates were paired in all possible combinations to determine their capacity for anastomosis. Mycelium plugs of isolates were placed 0.5 cm from each other on sterilized microscope slides covered with a thin layer of BM + NaNO3. Slides were then placed on petri dishes containing the same medium. The plates were incubated at 22ºC for 8 days. Afterwards, the mycelium plugs were removed from the slides; the newly grown hyphae were stained with cotton blue in lactophenol, coverslipped and examined under a light microscope. Obtaining heterokaryons and diploidsMycelium plugs (5 mm) of each nit mutant were paired equidistantly apart (approximately 1.0 cm) on petri dishes containing BM + NaNO3 for vegetative complementation tests. Dishes were incubated at 22ºC for 12 to 21 days and then examined for prototrophic heterokaryotic growth. Prototrophic (nit+) and homogeneous fast-growing sectors arising from heterokaryons grown in BM + NaNO3 were isolated and transferred to BM + NaNO3 + benomyl (0.5 µg/mL) to detect the presence of unstable diploids. Isolation of parasexual segregantsTwo methods were used to detect parasexual segregants (or sectors): 1) Isolation of sectors showing nit phenotype, directly from the heterokaryotic or prototrophic colonies. Mycelium plugs (5 mm) of these sectors were transferred to BM supplemented with different nitrogen sources for phenotypic characterization. 2) Suspensions of conidia were obtained from heterokaryons and diploid colonies growing in BM + NaNO3. Aliquots of these suspensions were later grown in BM + NaNO3 to recover recombinants and paternal segregants. Segregant ploidy was determined in BM + NaNO3 + benomyl (0.5 µg/mL): segregants that failed to produce mitotic sectors in the presence of the haploidization agent were classified as haploid segregants. Electrophoretic analysisMycelium plugs of nit mutants from R2047 and R23 races were individually grown in liquid CM. Cultures were kept at 22ºC, under shaking conditions, for five days. Mycelia obtained were blotted on filter paper after filtration and harvested individually. Two hundred milligrams mycelium from each strain was separately homogenized with a glass rod in an Eppendorf microcentrifuge tube using 80 µL 1.0 M phosphate buffer, pH 7.0, containing 5% PVP-40, 1.0 mM EDTA, 0.5% β-mercaptoethanol, 0.5% triton X-100, and 10% glycerol solution. After homogenization, the samples were centrifuged at 14,000 rpm for 30 min at 4ºC in a Beckman GS-15R centrifuge, and the supernatant (35 µL) was used from each sample. The polyacrylamide gels (15%) were prepared with 0.375 M Tris-HCl, pH 8.8, as buffer (Ceron et al., 1992). Electrophoresis was performed for 8 h at 4ºC, at a constant voltage of 200 V. Running buffer was 0.1 M Tris-glycine, pH 8.3. Staining techniques following a protocol established for plants (Pereira et al., 2001) identified the esterases. The gels were soaked for 30 min in 50 mL 0.1 M sodium phosphate, pH 6.2, at room temperature. Esterase activity was visualized by placing the gels for 1 h in a staining solution prepared from 50 mL sodium phosphate solution, 40 mg β-naphthyl acetate, 40 mg α-naphthyl acetate, 60 mg Fast Blue RR salt, and 5.0 mL N-propanol. Polyacrylamide gels were dried as described by Ceron et al. (1992) and Lapenta et al. (1995), and kept at room temperature for 1 h in a mixture of 7.5% acetic acid and 10% glycerol; the gels were then embedded in 5% gelatin, placed between two sheets of wet cellophane paper stretched on an embroidering hoop and left to dry for 24-48 h. The α-preferential esterases hydrolyzed preferentially α-naphthyl acetate in the presence of α- and β-naphthyl acetate and formed black bands; the β-preferential esterases hydrolyzed preferentially β-naphthyl acetate in the presence of α- and β-naphthyl acetate and formed red bands, and α/β-esterases hydrolyzed equally α- and β-naphthyl acetate, forming a mixture of black and red bands. The α-, β- and α/β-esterases were numbered in sequence, starting from the anode, according to decreasing negative charge. RESULTS Isolation of nitrate non-utilizing mutantsStable mutants obtained in the presence of potassium chlorate, which produced poor growth colonies, or rather, without dense aerial mycelium, were considered mutants incapable of using nitrate (nit). Mutants were screened for their ability to utilize different nitrogen sources in order to determine their phenotypes and could be divided into three distinct phenotypic classes: nit1 (nitrate non-utilizing), Nit3 (nitrite and nitrate non-utilizing), and NitM (hypoxanthine and nitrate non-utilizing). One hundred and twenty-eight nitrate non-utilizing mutants were obtained from the five C. lindemuthianum isolates. Seventy-six (59%) were classified as nit1 mutants, nineteen (15%) were characterized as Nit3 mutants, and thirty-three (26%) typified NitM mutants, utilizing nitrite but not hypoxanthine (results not shown). The remaining mutants, which showed either phenotypic instability when grown in BM + NaNO3 or could utilize nitrate, were discarded. These unstable mutants were characterized as either heterokaryons containing both nit and nit+ nuclei, or containing a distinct mutation conferring both nitrate-utilizing and chlorate-resistance phenotypes. Anastomoses between isolates and heterokaryon formationAll isolates were tested for anastomosis and heterokaryon formation. Isolates R2047, R65 and R23 were found to carry out anastomosis among themselves but none of them formed anastomoses with the R73 isolate. R89 isolate, on the other hand, was capable of carrying out anastomosis with all the isolate ones, except with R2047 (Figure 1A). 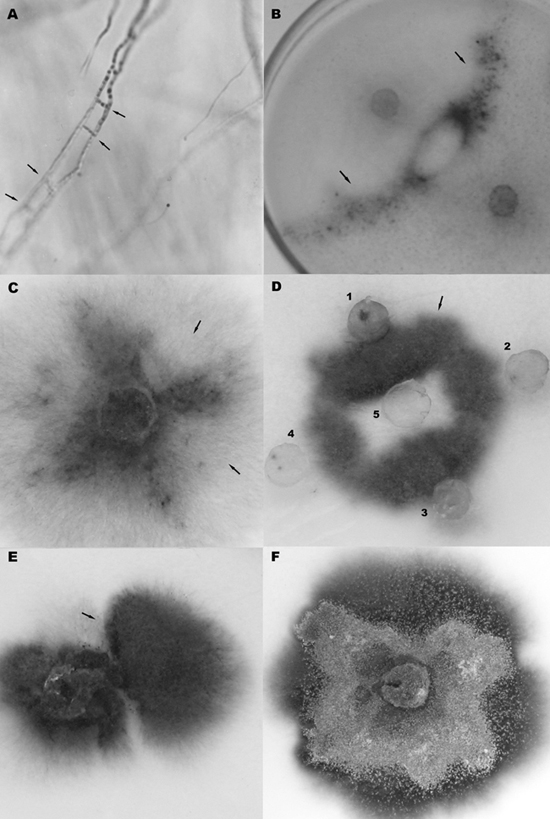
Figure 1. A. Hyphal anastomoses (arrows) among nit mutants of R2047 and R23 isolates. B. Pairing between NitM mutants from R65 and R23 races separated by a dialysis membrane. Heterokaryons (arrows) did not occur beneath the dialysis membrane which impairs direct contact between the strains’ mycelia. C. Heterokaryotic colony showing mitotic segregation of the nit paternal mutants (arrows). D. Heterokaryons (arrow) formed between NitM mutants of R65 (1, 3, 4), R23 (2), and R2047 (5). E. Heterokaryon colony showing a fast-growing sector (arrow). F. Unstable diploid strain growing in BM + benomyl. Pairing of complementary nit mutants produced colonies of prototrophic growth in BM + NaNO3, after two or three weeks of incubation at 22ºC. Since no heterokaryotic growth occurred when the paired mutant strains were separated by a dialysis membrane (Figure 1B), this fact indicated that the prototrophic growth is not due to cross-feeding. Prototrophic colonies gave rise spontaneously to auxotrophic sectors (or parasexual segregants) in BM + NaNO3, which showed parental (i.e., nit1, Nit3 or NitM) phenotypes (Table 1, Figure 1C). Results indicate that the prototrophic growth from the pairing of different complementary auxotrophic mutants must be due to heterokaryosis. To determine whether the isolates were vegetatively self-compatible, nit1 mutants were paired with either Nit3 or NitM mutants from the same strain on BM + NaNO3. Mutants were seen to be self-compatible due to the formation of dense heterokaryons (results not shown). When pairing the complementary nit mutants of R2047, R65 and R23 isolates in all possible combinations, interstrain barriers of vegetative incompatibility were not observed. Therefore, isolates R2047, R23 and R65 were allocated in the vegetative compatibility group I (VCG I). In addition, mutants NitM of R2047, R65 and R23 races showed vegetative complementation among themselves and formed a dense line of growth in the contact area between the colonies (Table 1, Figure 1D). On the other hand, pairings R2047/R89, R2047/R73, R73/R65, and R73/R23 showed complete vegetative incompatibility by this method (results not shown), and R73 isolate was allocated in another vegetative compatibility group (VCG II). Diploid formation and characterizationFast-growing sectors arise spontaneously from prototrophic heterokaryotic colonies two or three weeks after pairing in BM + NaNO3 (Figure 1E). Sectors showed nit+ phenotype and growth rate similar to that of the original wild-type strains in BM + NaNO3. When conidia or mycelium plugs from prototrophic sectors were transferred to BM + NaNO3 or benomyl-supplemented BM + NaNO3, their mitotic instability could be identified by the production of auxotrophic segregants (or sectors), which showed parental (NitM or nit1) phenotypes (Table 1, Figure 1F). The results suggest that fast-growing sectors are at least transiently diploid. The diploid strains R2047.4 (NitM) // R23.16 (NitM) and R2047.4 (NitM) // R23.19 (NitM), and the parental haploid strains, R2047.4, R23.16 and R23.19, were characterized for esterase isoenzyme expression, and differential α- and β-esterase patterns were observed (Figure 2). A total of 27 α- and β-esterases were identified, but EST-7, EST-13, EST-17, and EST-22 were present only in the R2047.4 haploid strain. EST-6, EST-8, EST-14, EST-16, EST-18, EST-20, EST-23, EST-24, EST-25, and EST-26 isoenzymes were detected only in the R23.16 and R23.19 strains. Strain-specific esterases indicate specific biochemical characteristics, and differential expression of esterase loci have in fact been demonstrated in the filamentous fungus Aspergillus nidulans (Machado and de Castro-Prado, 2001). On the other hand, in diploid strain R2047.4 // R23.19, both paternal and additional (EST-9, EST-10, EST-12, EST-14, and EST-27) isoenzyme esterases were detected (sample 3, Figure 2). Figure 2. Polyacrylamide gel electrophoresis showing α- and β-esterases in haploid strains R2047.4 (sample 1), R23.19 (sample 2), and R23.16 (sample 4), and in diploid strains R2047.4 (NitM) // R23.19 (NitM) (sample 3), and R2047.4 (NitM) // R23.16 (NitM) (sample 5). The Est-7, Est-13, Est-17, and Est-22 isozymes were present only in the R2047.4 haploid strain, whereas Est-6, Est-8, Est-14, Est-16, Est-18, Est-20, Est-23, Est-24, Est-25, and Est-26 isozymes were detected only in the R23.16 and R23.19 strains. DISCUSSION Nitrate non-utilizing mutants were used to demonstrate heterokaryosis and parasexual segregation in C. lindemuthianum isolates. Auxotrophic complementation and segregation of nutritional markers have also been used to characterize parasexuality in M. grisea and F. moniliforme (Crawford et al., 1986; Klittich and Leslie, 1988). Heterokaryon formation by the anastomosis of homokaryotic hyphae contributes to the high degree of genetic variability encountered in filamentous fungi, and provides a mechanism for the restoration of normal function in combined, separately deficient, genomes (Zeigler, 1998; Saupe, 2000). Contrastingly, somatic incompatibility, which impairs heterokaryon formation among different mycelia, may affect the parasexual occurrence (Saupe, 2000). In the present study, races R2047, R65 and R23 were shown to integrate the VCG I, whereas the R73 isolate was allocated in VCG II. Vegetative incompatibility between R2047 and R73 isolates were also reported by Roca et al. (2004). On the other hand, nit mutants from R89 isolate were able to carry out anastomosis and to form heterokaryons with nit mutants of isolates from both VCG I (R65 and R23) and VCG II (R73). The R89 isolate may be characterized as a “bridge” strain, analogous to the C. lindemuthianum Mi.7.99 isolate (Rodríguez-Guerra et al., 2003), which was capable of carrying out anastomosis with isolates from two anastomosis groups. In Neurospora crassa and Fusarium species, multiple loci control the production of a molybdenum-containing co-factor (Tomsett and Garrett, 1980; Klittich and Leslie, 1988). Prototrophic recombinants were in fact recovered from meiotic crosses of nitM mutants of F. moniliforme strains of opposite mating type (Klittich and Leslie, 1988). The vegetative complementation of nit mutants with the same phenotype shown in this study (Figure 1D, Table 1), also suggests the existence of multiple loci that control the production of the molybdenum-containing co-factor in C. lindemuthianum. Further studies will be necessary to determine how many loci are involved in the production of the NitM phenotypes. The association of genetically dissimilar nuclei in heterokaryotic mycelia may form stable heterozygous diploid nuclei through the fusion of two complete genomes. Diploid colonies can be distinguished easily from haploids on the basis of their restrictive growth and breakdown in the presence of benomyl, the haploidization agent. Benomyl may produce haploidy via successive chromosome losses and mitotic non-disjunction (Pontecorvo et al., 1953; Hastie, 1970; Miyamoto et al., 2007). In the present study, diploid strains were obtained spontaneously from heterokaryons formed between complementary nit mutants. The results of haploidization of the diploid strains, presented in Table 1, demonstrate that heterokaryons in C. lindemuthianum may yield unstable diploid conidia. Although all diploid strains gave rise to haploid paternal segregants, the phenotypic analyses of these segregants demonstrated that certain parental genotypes were not recovered or recovered only at low frequency. The differential expression of esterase loci in diploid and respective haploid paternal strains is more an evidence of the occurrence of parasexuality in the C. lindemuthianum pathogen, underpinning the fact that different forms of nucleosome organization in chromatin may increase or repress the expression of specific genes (Lee and Young, 2000), or that recombination events in diploid strains may generate differential α- and β-esterase phenotypes. The present study provides evidence of both heterokaryosis and diploid formation in C. lindemuthianum under laboratory conditions, using nitrate non-utilizing mutants. REFERENCES Baayen RP, Forch MG, Waalwijk C, Bonants PJM, et al. (1998). Pathogenic, genetic and molecular characterization of Fusarium oxysporium f. sp. Lilii. J. Plant Pathol. 104: 887-894. Balardin RS, Jarosz AM and Kelly JD (1997). Virulence and molecular diversity in Colletotrichum lindemuthianum from South, Central and North America. Phytopathology 87: 1184-1191. Ceron CR, Santos JR and Campos Bicudo HEM (1992). The use of gelatin to dry cellophane wound slab gels in an embroidering hoop. Braz. J. Genet. 15: 201-203. Correl JC, Harp TL, Guerber JC, Zeigler RS, et al. (2000). Characterization of Pyricularia grisea in the United States using independent genetic and molecular markers. Phytopathology 90: 1396-1404. Crawford MS, Chumley FG, Weaver CG and Valent B (1986). Characterization of the heterokaryotic and vegetative diploid phases of Magnaporthe grisea. Genetics 114: 1111-1129. Hastie AC (1970). Benlate-induced instability of Aspergillus diploids. Nature 226: 771. Jerba VF, Rodella RA and Furtado EL (2005). Relationship between bean leaf structure and the Glomerella cingulata f.sp. phaseoli preinfection. Pesqui. Agropecu. Bras. 40: 217-223. Kimati H and Galli F (1970). Glomerella cingulata f. sp. phaseoli, fase ascógena do agente causal da anthracnose do feijoeiro. An. Esc. Sup. Agric. Luiz de Queiroz 27: 411-437. Klittich C and Leslie JF (1988). Nitrate reduction mutants of Fusarium moniliforme (Gibberella Fujikuroi). Genetics 118: 417-423. Lapenta AS, de Campos Bicudo HE, Ceron CR and Cordeiro JA (1995). Esterase patterns of species in the Drosophila buzzatii cluster. Cytobios 84: 13-29. Lee TI and Young RA (2000). Transcription of eukaryotic protein-coding genes. Annu. Rev. Genet. 34: 77-137. Machado MF and de Castro-Prado MA (2001). Differential esterase expression in developmental mutants of Aspergillus nidulans. Biochem. Genet. 39: 357-368. Mahuku GS and Riascos JJ (2004). Virulence and molecular diversity within Colletotrichum lindemuthianum isolates from Andean and Mesoamerican bean varieties and regions. Eur. J. Plant Pathol. 110: 253-263. Miyamoto CT, Sant’Anna JR, Franco CC and Castro-Prado MA (2007). Genotoxicity (mitotic recombination) of the cancer chemotherapeutic agents cisplatin and cytosine arabinoside in Aspergillus nidulans. Food Chem. Toxicol. 45: 1091-1095. Pereira AJ, Lapenta AS, Vidigal-Filho PS and Machado MF (2001). Differential esterase expression in leaves of Manihot esculenta Crantz infected with Xanthomonas axonopodis pv. manihotis. Biochem. Genet. 39: 289-296. Pontecorvo G, Roper JA, Hemmons LM, Macdonald KD, et al. (1953). The genetics of Aspergillus nidulans. Adv. Genet. 5: 141-238. Roca MG, Davide LC, Mendes-Costa MC and Wheals A (2003). Conidial anastomosis tubes in Colletotrichum. Fungal Genet. Biol. 40: 138-145. Roca MG, Davide LC, Davide LM, Mendes-Costa MC, et al. (2004). Conidial anastomosis fusion between Colletotrichum species. Mycol. Res. 108: 1320-1326. Rodríguez-Guerra R, Ramírez-Rueda MT, De la Veja OM and Simpson J (2003). Variation in genotype, pathotype and anastomosis groups of Colletotrichum lindemuthianum isolates from México. Plant Pathol. 52: 228-235. Saupe SJ (2000). Molecular genetics of heterokaryon incompatibility in filamentous ascomycetes. Microbiol. Mol. Biol. Rev. 64: 489-502. Tomsett AB and Garrett RH (1980). The isolation and characterization of mutants defective in nitrate assimilation in Neurospora crassa. Genetics 95: 649-660. Zeigler RS (1998). Recombination in Magnaporthe grisea. Annu. Rev. Phytopathol. 36: 249-275. |
|