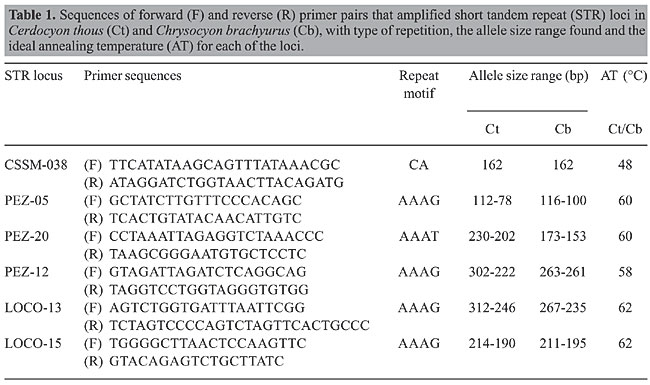
ABSTRACT. The maned wolf (Chrysocyon brachyurus) and the crab-eating fox (Cerdocyon thous) are two wild-canid species found in the Brazilian Cerrado. We tested cross-amplification and transferability of 29 short tandem repeat primers originally developed for cattle and domestic dogs and cats on 38 individuals of each of these two species, collected in the Emas National Park, which is the largest national park in the Cerrado region. Six of these primers were successfully transferred (CSSM-038, PEZ-05, PEZ-12, LOCO-13, LOCO-15, and PEZ-20); five of which were found to be polymorphic. Genetic parameter values (number of alleles per locus, observed and expected heterozygosities, and fixation indices) were within the expected range reported for canid populations worldwide. Key words: STR markers, Cross-amplification, Cerdocyon thous, Chrysocyon brachyurus, Cerrado The ‘Emas’ National Park, situated in south-western Goiás State, Central Brazil, is currently the largest national park in the Cerrado; it possesses a very representative fauna, including most of the large-bodied mammals that inhabit this ecosystem within Brazil (Marinho-Filho et al., 2002; Rodrigues et al., 2002). Understanding the population genetic structure of species living in these regions is important because the entire Cerrado region is now under extreme threat (Myers et al., 2000) and thus the large-bodied mammals are expected to be preserved mostly within the few remaining large conservation units. It is clear that conserving native species and understanding their population processes require knowledge about their genetic diversity (Hedrick, 2001). Microsatellite markers, or short tandem repeats (STR), are now considered the best choice to obtain genetic data, with a good balance between costs, information context and operational facilities; they have become a useful tool for population and conservation genetic studies (Li et al., 2002; Brondani et al., 2003; Wandeler et al., 2003). However, STR markers are usually developed for each species, which is sometimes difficult and involves considerable laboratory work. Another strategy is to look for cross-amplification of markers among phylogenetically related species for population and conservation genetic analyses (e.g., Tong et al., 2002; Williamson et al., 2002; Zucchi et al., 2002; Klukowska et al., 2003). We tested the transferability of STR markers originally developed for domestic cattle, buffaloes and domestic dogs and cats for two species of Canids found in the Emas National Park, the crab-eating fox, Cerdocyon thous, and the maned wolf, Chrysocyon brachyurus. The maned wolf is the largest canid in South America, with important ecological roles as a top-chain predator and seed disperser; while the crab-eating fox is a relatively small-bodied, abundant and widely distributed species, found in many different habitats and biomes of Brazil. Genomic DNA was extracted from blood of 38 individuals of each species captured in the Emas National Park. We tested 29 pairs of primers, initially using only two individuals of each species (a complete list of the primers tested is available from the authors upon request). Among these primers, six pairs were successfully transferred to the target species; these were then used for genotyping the 38 individuals of each species (Table 1).
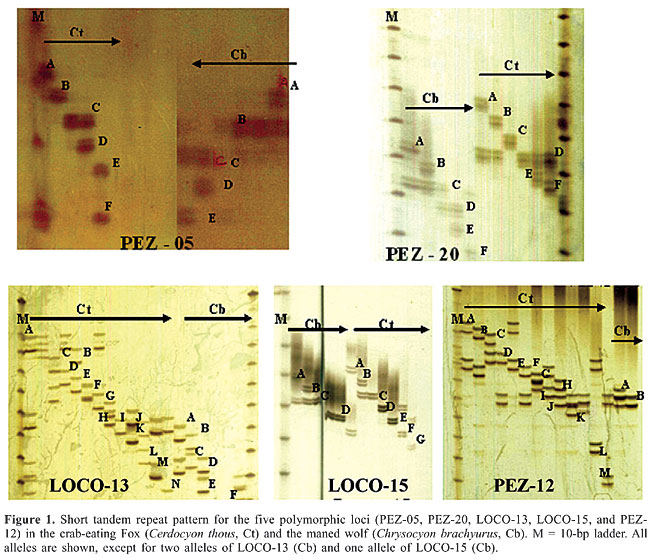
The PCR reactions were done in a final volume of 15 µL, containing 5 µL DNA (3 ng/µL), 2.2 µL of each of the primers (0.9 mM), 1.3 µL bovine serum albumin (10 mg/mL), 1.5 µL PCR buffer (10X), 0.5 µL MgCl2 solution (50 mM), 1.3 µL d’NTP solution (2.5 mM), and one unit of Taq DNA Polymerase (furnished by Amersham Pharmacia Biotech). The reactions were run in a 96-well PT-100 thermocycler (MJ Research), with an initial 5-min incubation at 94°C, followed by 30 cycles at 94°C for 1 min, annealing of “primers” from 45 to 65°C, and a final extension at 72°C for 7 min. The amplified products were electrophoresed on 4% denaturing acrylamide gels running in 1X TBE at 90 W for 2 h, and then stained with silver nitrate (Bassam et al., 1991). DNA ladders (standard fragment size 10 bp - Amersham Pharmacia Biotech) were used to estimate the size of each fragment. The primer pairs were classified according to the quality of amplified PCR products obtained. About half of the pairs of primers (14 pairs) did not reveal any amplified products, whereas nine pairs gave non-specific amplifications. The remaining six pairs were successfully transferred to the two species. These six primers were originally developed for the domestic dog (PEZ-05, PEZ-12, LOCO-13, LOCO-15, and PEZ-20) (Halverson et al., 1999) and cattle (CSSM-038) (Barker et al., 1997). We then analyzed the patterns for 38 individuals of each species; five of six loci showed polymorphism. We found on average 7.8 alleles per locus in the crab-eating fox and 4.5 alleles per locus in the maned wolf at the polymorphic loci (Figure 1).
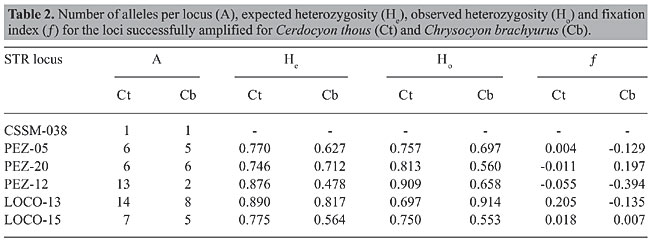
Based on these five polymorphic loci, we obtained initial estimates of some genetic parameters in these two species. The average expected heterozigosity (He) for the crab-eating fox was 0.676, whereas the observed heterozigosity (Ho) was 0.654, resulting in a fixation index (¦) of 0.032. The parameters for the maned wolf were He = 0.533, Ho = 0.564, and ¦ = -0.091 (Table 2). These values are within the expected range of values, considering previous knowledge of canid population genetics worldwide (Wayne, 1996).
We confirmed that there is great potential for cross-amplification of STR markers between different canid species. This is important for genetic studies of Neotropical canids, and mammals in general, because expensive and time-consuming development of species-specific primers is not strictly necessary for initiating population and conservation genetic studies in these species. This type of polymorphism data is considered critical for understanding population processes and to establish optimal management strategies. ACKNOWLEDGMENTS Research supported by the PRONEX program of CNPq and SECTEC-GO (Proc. 23234156). J.A.F. Diniz-Filho was partially supported by a CNPq 1A productivity fellowship, whereas M.P.C. Telles is the recipient of a PROPE/UCG fellowship. REFERENCES Barker JS, Moore SS, Hetzel DJ, Evans D, et al. (1997). Genetic diversity of Asian water buffalo (Bubalus bubalis): microsatellite variation and a comparison with protein-coding loci. Anim. Genet. 28: 103-115. Bassam BJ, Caetano-Anolles G and Gresshoff PM (1991). Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal. Biochem. 196: 80-83. Brondani C, Rangel PH, Borba TC and Brondani RP (2003). Transferability of microsatellite and sequence tagged site markers in Oryza species. Hereditas 138: 187-192. Halverson J, Dvorak J and Stevenson T (1999). Microsatellite sequences for canine genotyping. US Patent 05874217. Hedrick PW (2001). Conservation genetics: where are we now? Trends Ecol. Evol. 16: 629-636. Klukowska BJ, Strabel T, Mackowski M and Switonski M (2003). Microsatellite polymorphism and genetic distances between the dog, red fox and artic fox. J. Anim. Breed. Genet. 120: 88-94. Li YC, Korol AB, Fahima T, Beiles A, et al. (2002). Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review. Mol. Ecol. 11: 2453-2465. Marinho-Filho JS, Rodrigues FH and Juarez KM (2002). The cerrado mammals: diversity, ecology and natural history. In: The cerrados of Brazil: ecology and natural history of a Neotropical savanna (Oliveira PS and Marquis RJ, eds.). Columbia University Press, Irvington, 266-284. Myers N, Mittermeier RA, Mittermeier CG, da Fonseca GA, et al. (2000). Biodiversity hotspots for conservation priorities. Nature 403: 853-858. Rodrigues FH, Silveira L, Jácomo AT, Bezerra AP, et al. (2002). Composição e caracterização da fauna de mamíferos do Parque Nacional das Emas, Goiás. Rev. Bras. Zool. 19: 589-600. Tong J, Wang Z, Yu X, Wu Q, et al. (2002). Cross-species amplification in silver carp and bighead carp with microsatellite primers of common carp. Mol. Ecol. Notes 2: 245-247. Wandeler P, Funk SM, Largiader CR, Gloor S, et al. (2003). The city-fox phenomenon: genetic consequences of a recent colonization of urban habitat. Mol. Ecol. 12: 647-656. Wayne RK (1996). Conservation genetics in the Canidae. In: Conservation genetics: case histories from nature (Avise JC and Hanuick JL, eds.). Chapman and Hall, New York, 75-118. Williamson KS, Cordes JF and May B (2002). Characterization of microsatellite loci in Chinook salmon (Oncorhynchus tshawytscha) and cross-species amplification in other salmonids. Mol. Ecol. Notes 2: 17-19. Zucchi MI, Brondani RP, Pinheiro JB, Brondani C, et al. (2002). Transferability of microsatellite markers from Eucalyptus spp. to Eugenia dysenterica (Myrtaceae family). Mol. Ecol. Notes 2: 512-513. |
|